This blog post by Jennifer Glass, assistant professor in the School of Earth and Atmospheric Sciences at Georgia Institute of Technology, chronicles her journey from first falling in love with marine science as a college freshman, to the first original research paper out of her own lab, now openly accessible:
My first scientific loves were the ocean and metals, in that order. From the window of my freshmen college dorm room, I could see Monterey Bay through the redwoods. In the summer, upwelling of deep nutrients to the surface caused phytoplankton blooms that turned the whole surface of the bay a visible cyan. When I learned the mechanism behind that color change, I knew I wanted to study science; it explained how minuscule things like single-celled bacteria and algae could change the face of the entire Earth.
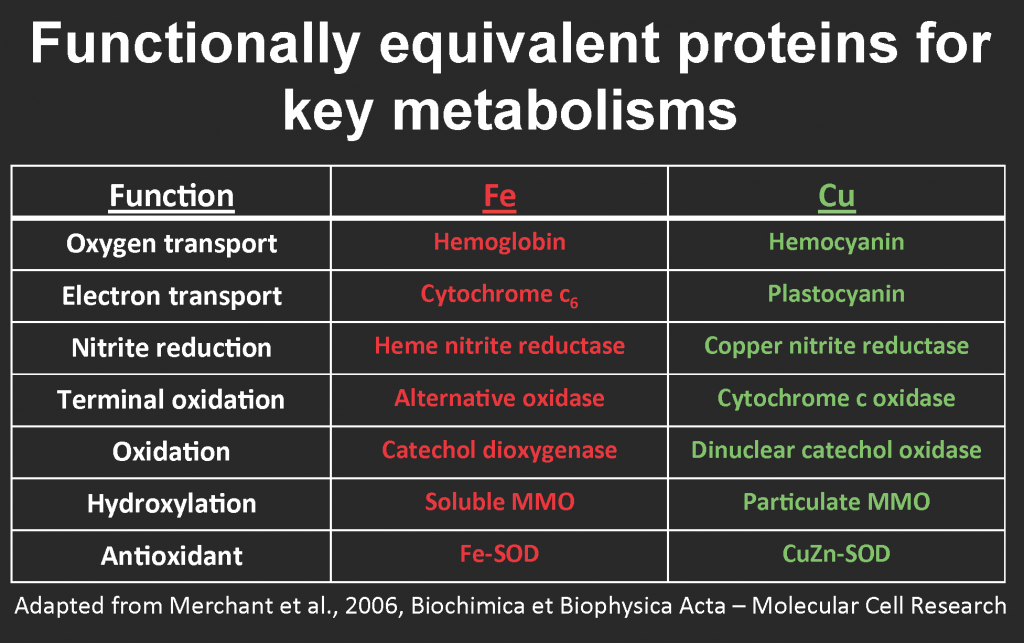
Blood. Tiny tweaks in the structure of porphyrin rings and the nature of their metal center produce a broad diversity of bioessential molecules, including the red iron-containing hemoglobin that carries oxygen in animal blood, the blue copper-containing hemocyanin that transports oxygen in mollusks, and the green magnesium-containing chlorophyll that plants use to harvest light energy. A simple explanation for the utilization of different metals is availability: most land animals have access to plenty of iron in soils, whereas some invertebrates apparently swapped it out for more soluble copper in oxygenated waters. Fascinatingly, clams and worms at deep-sea iron-rich hydrothermal vents also use hemoglobin, unlike their iron-starved brethren.
Hemoglobin/hemocyanin is only one example of iron-copper swapping. A similar “biochemical switch” also evolved in open-ocean diatoms that substitute copper-containing plastocyanin for functionally equivalent iron-containing cytochrome c6 in the photosynthetic electron transport, enabling them to ration their iron reserves for other vital metabolic needs (Peers & Price, 2006). The same swap occurs in copper-starved cultures of the green alga Chlamydomonas reinhardtii (Merchant et al. 2006). Some aerobic methanotrophs can substitute copper for iron in a variant of the methane monooxygenase enzyme (reviewed by Semrau et al., 2010; for more info, see Glass 2015 and table below).
It seems likely that the evolution of iron proteins preceded those containing copper (Dupont et al., 2010). Before the rise of atmospheric oxygen some 2.4 billion years ago, the ocean was rich in dissolved iron. That evolutionary fingerprint remains in extant life despite the scant availability of iron in modern earth’s surface ecosystems. In other words, in contrast to human history, the Iron Age came before the Bronze Age in earth history.
Sweat. I wondered whether we might also find evidence for marine microbes meddling with iron and copper in modern seawater today. I had the opportunity to test this hypothesis on a 2013 research cruise to the Eastern Tropical North Pacific oxygen minimum zone (OMZ), led by my colleague Dr. Frank Stewart, assistant professor at Georgia Tech. This cruise was later christened “OMZoMBiE” (“Oxygen Minimum Zone Microbial Biogeochemistry Expedition”) to reflect the scarcity of sleeping hours.
A video compilation of cruise photos is shown below; the soundtrack was selected in recognition of the participation of both Dr. Alejandro Murillo from Universidad de Concepción, Chile, and Dr. Alejandra Prieto Davó from Universidad Nacional Autónoma de México, as well as the apt Lady Gaga lyric hot like Mexico, rejoice (the west coast off Manzanillo is indeed toasty in June).
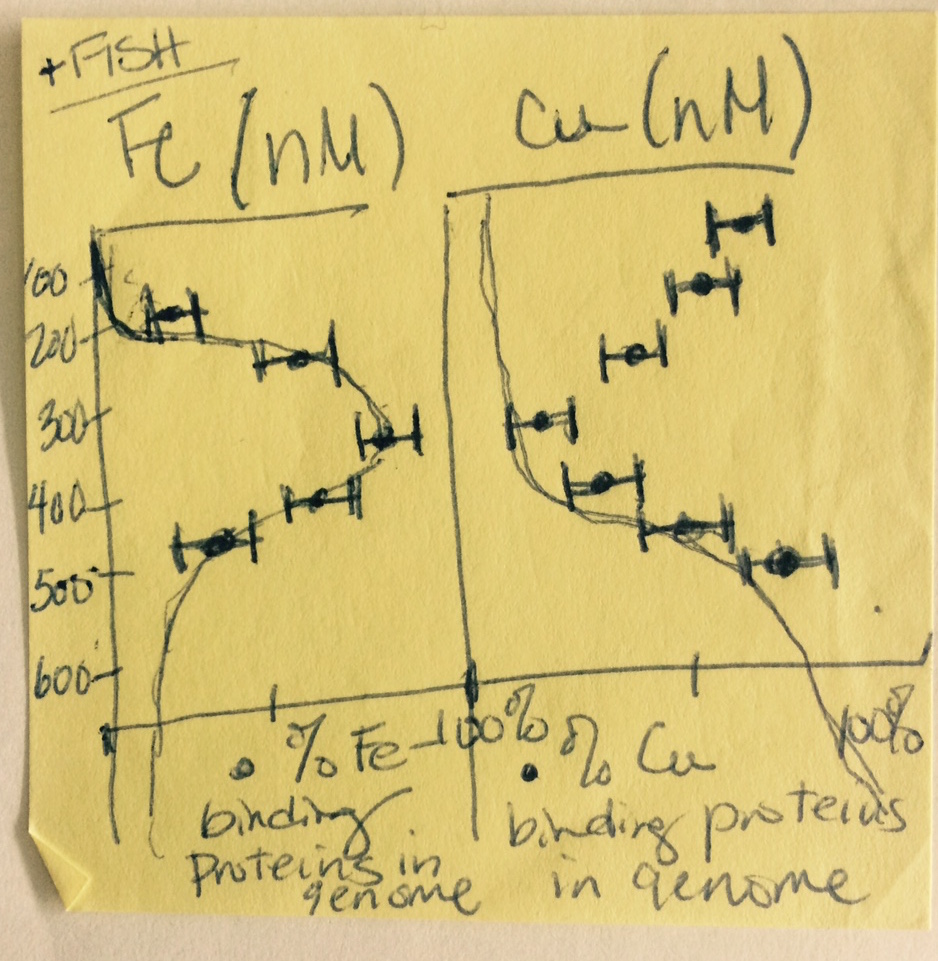
My goal for the cruise was to track correlations between genes and transcripts encoding metalloenzymes and marine metal availability via careful measurement of the trace metal content of the same seawater samples that would be sequenced for DNA and RNA. Below is a quick sketch I made on a Post-It note the first night aboard the New Horizon research vessel. It shows the general trends in depth profiles I expected to find. (The note at the top “+FISH” was to remind myself to also take samples for Fluorescence In Situ Hybridization, not to actually go fishing!)
In order to collect the samples to get these data, I first had to acid wash 160 plastic bottles by hand with 38 liters of hydrochloric acid, acquire all-plastic “Go-Flo” Niskin water sampling bottles (kindly lent by supportive colleagues Kathy Barbeau and James Moffett) and construct an all-plastic trace metal clean mini “bubble” lab that was constantly inflated with filtered air to protect my clean seawater samples from particle contamination that was everywhere on the slowly rusting ship. A helpful “cookbook” for this sampling technique is available online, and many people assisted me in these efforts (see the acknowledgements sections of the paper for all of their names).
Tears. The research cruise was filled with long days and equally long nights of water sampling and filtering. During one particularly heartbreaking 3 am shift, we accidentally deployed the Go-Flo bottle to 500 meters water depth before it had equilibrated with the greater pressure of the overlying water mass. The top seal popped off, rendering it useless for the remainder of the cruise and requiring several thousands of dollars worth of repairs to be paid out of my precious start-up funds.
After opening my own lab that autumn, I learned that science, even with the most careful planning and oversight, is expensive, and frequently and inevitably fraught with failure. “I write this paragraph with ease,” wrote Professor Erwin F. Smith – an early plant pathologist and the first to show that bacteria can cause plant disease – in his “Report on Peach Yellows” (Bulletin 9 of the 1889 USDA Botanical Division),
“….but the work is full of difficulties. Nature does not yield her secrets upon the mere asking. Only those engaged in similar inquiries can have any adequate conception of the labor involved or of the perplexities which can beset one at every step. Moreover, in such an inquiry nothing can be promised in advance. The investigator and the public alike must take their chances on the results. However… there seems to be every encouragement for the renewed and persistent prosecution of this inquiry. By such effort sources of error will be discovered, difficulties overcome and the truth finally discovered.”
Or, as astrophysicist Sarah Tuttle put it in her viral tweet 125 years later: “Failure required, tears optional. Please don’t lick the science.”
Ok. I made a lab sign, if anyone needs one. #TimHunt #DistractinglySexy #WomenInScience #STEM #SafeLab pic.twitter.com/teezqx4rO5
– Sarah Tuttle (@niais) June 13, 2015
Now, more than two years after that hot June in Mexico, the datasets are finally complete and my lab’s first original research paper is finally published. The trends, while not as perfect as I had sketched the first night of the cruise, generally held as predicted. They are also the first environmental community-level evidence for higher relative abundance of genes encoding iron-containing proteins in an oxygen-starved ecosystem, with the opposite trend for genes encoding copper-containing proteins, namely cytochrome c oxidase, in oxygen-rich waters.
Several hundred meters beneath the ocean surface off the west coasts of Mexico, Chile, Namibia and India, the Iron Age of earth history still reigns supreme. These OMZs are a fossil relict of what the entire ocean might have looked like millennia before the first oxygen-generating cyanobacteria changed Earth’s face forever. Slowly, they are revealing their secrets.
Contact Form to Provide Feedback Directly to Jennifer Glass About This Article