Starting in the fall of 2012 we began a second project to involve undergraduates (and a high school student) in real research that relates to the microbiology of the built environment. This time we looked at biogeography and succession in microbial communities found in aquariums. A description of the project can be found here and the project blog is here. With the end of the academic year, the project has officially wrapped up as far as the students are concerned but there is much more work to be done.
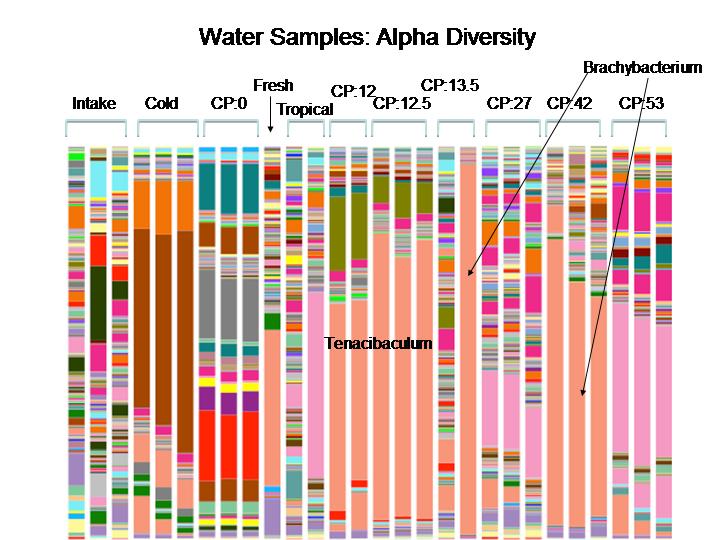
For our first pass we extracted DNA and make 16S libraries from about 100 samples taken from the water, sediment, and walls of a newly established aquarium tank over time, as well as some other environments for reference. After demultiplexing the barcoded libraries we began to play around with the data in QIIME. As it turned out, this was a whole project in itself… particularly for the students with no experience using the command line. We have some interesting preliminary data and are currently in the process of prepping an additional ~200 samples for sequencing. Once we have all the data in hand we’ll begun the full analysis in earnest. For a taste of the data, check out this graph showing the alpha diversity between the various water samples. There will be much much more like this once we finish sequencing the additional samples and pool all the data for analysis. Stay tuned!
New at #microBEnet: Progress Report on Undergraduate Aquarium Biogeography Project: Starting in the fall of 20… http://t.co/mrXNrIVp7l
RT @Dr_Bik: New at #microBEnet: Progress Report on Undergraduate Aquarium Biogeography Project: Starting in the fall of 20… http://t.co/m…
Microbes: Progress Report on Undergraduate Aquarium Biogeography Project http://t.co/8kqTSljJ1k via @microBEnet #micrBE.net
RT @Dr_Bik: New at #microBEnet: Progress Report on Undergraduate Aquarium Biogeography Project: Starting in the fall of 20… http://t.co/m…
Progress Report on my labs Undergraduate Aquarium Biogeography Project http://t.co/rmlcR5zSbW via @microBEnet
RT @phylogenomics: Progress Report on my labs Undergraduate Aquarium Biogeography Project http://t.co/rmlcR5zSbW via @microBEnet
RT @phylogenomics: Progress Report on my labs Undergraduate Aquarium Biogeography Project http://t.co/rmlcR5zSbW via @microBEnet