(This blog post was prepared by students enrolled in the Koala Poop Microbiome Class in the Fall of 2016 at UC Davis)
This week we set out to extract DNA from isolated bacteria grown in liquid culture to further analyze it. Students in the lab were assigned test tubes containing tannin degrading bacteria which have been isolated and were in charge of extracting the bacterial DNA. Using the Qiagen DNeasy Blood and Tissue Kit, we separated DNA with ethanol, that caused the DNA to form a solid and precipitate out of the solution. Fun Fact: The precipitation of DNA using ethanol has a lot more to do with physics that you might think click on the link to see http://physiology.med.cornell.edu/faculty/mason/lab/zumbo/files/ETHANOL_PRECIPITATION.pdf) Our kit uses spin columns.
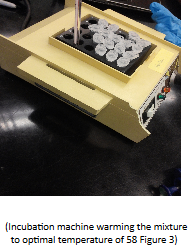

We decided that DNA extractions are a tedious process. We hope our isolated DNA could be from a bug that’s interesting to the science community. This class is always busy; each lab session is full of new and exciting learning opportunities. Students work separately and as part of a group effort. In this week’s lab everyone works on their own extractions but also in collaboration as a team. Group shared pipets, took turns to using chemical reagents, and compared their results upon completion of each step. Working in a lab has its benefits, I learned this when I made random errors with my buffer. A teammate pointed out the various buffers and that our solutions appeared different despite being at the same step. However, this didn’t discourage me to continue as I was able to work with a classmate and help him finish his extractions. It is true that sometimes mistakes give rise to answers, and that not all that hard work it’s lost when mistakes do happen. Now for the science community, what is your consideration of an efficient and organized way of going about this method of DNA extraction to reduce mistakes?
Next week we will be amplifying DNA collected in this week’s lab. This will give many students the opportunity to have hands on experience with PCR (polymerase chain reaction). Hopefully, with the success of this lab we can further the progress on the class project.