(This blog post was prepared by students enrolled in the Koala Poop Microbiome Class in the Fall of 2016 at UC Davis)
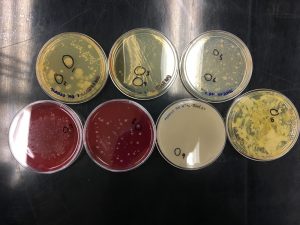
The goal of this week was to isolate a single bacterial colony for further study. This week we observed bacterial growth from our koala poop samples. With a partner, we obtained and observed the individual plate cultures from Week 1 to see if anything had grown. We then chose and labeled 10 colonies, particularly ones that were large in size and clearly isolated from other colonies. In order to start the process of dilution streaking, we first obtained 10 new agar plates. Each pair made sure that these new agar plates had the same media type as Week 1.
We hope to identify the kinds of plates that the isolates were most prevalent on, and maybe how media relates to the koala’s diet. With this isolated bacteria, we will be able to carry on with the experiment and extract DNA (Week 3!). Click here to learn more information about dilution streaking.
It was interesting to find several different kinds of bacteria in the petri dishes prepared last week. Looking around the lab, we could see tannin-degrading bacterial colonies that were on many of the plates ! The dilution streaking was a little challenging since we had to switch inoculating loops to do it properly, and the agar could be easily punctured up with the loops. Keeping track of the plates used was confusing because some of us not only used our own plates, but also used plates from the Australian collection in order to get the ten colonies required per group.
In preparation for Week 3, the class also practiced using micropipettes. In table groups, we learned to efficiently pipette small amounts of water in and out of an eppendorf tube as part of a game. This was somewhat challenging because not everyone at our table had the same micropipette so each dial was labelled differently, however, it was a fun experience! Some questions coming into Week 3 are: What kind of bacteria colonies were on the plates? What bacteria was the most common among all the plates? Are there other methods similar to streak plating that isolate microorganisms?