We’re recruiting a student/postdoc for this project! If it sounds interesting, please contact the Huttenhower Lab!
We were happy to hear that we’ve been funded by the Sloan Foundation to continue our study of microbes on the Boston subway. Our original study involved 1) identifying which microbes were resident in this built environment, 2) understanding their functional potential, and 3) quantifying gene families of interest, including antibiotic resistance and virulence factors. In brief, we found that it’s not so bad – even (and perhaps especially) on a crowded subway, you’re surrounded mainly by harmless microbes largely from human skin.
However, this still leaves many open questions in terms of public health. Although we now know what microbes are present, the built-environment is chemically unique, and we have yet to characterize which microbes are biochemically active, persistent, or dead. Metagenomic surveys reveal which metabolic pathways are present, but this may include DNA from inactive cells. It also does not identify which pathways are transcriptionally active, driving the cells that do persist in the built environment. Finally, since we collected all train samples at one time point and all touchscreen samples at another time point, we have limited knowledge on how stable different locations’ microbial activities are over time.
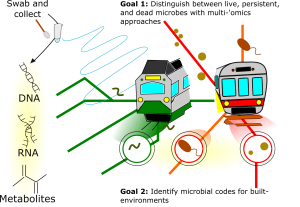 To determine which microbes are active, we will use a combination of ‘omics approaches on newly collected samples that include metagenomics, metatranscriptomics, and metabolomics. From this, we can identify live bacteria (DNA) based on those that are transcriptionally active (RNA). We can further determine which metabolites are being produced by 1) associating taxonomic profiles with metabolites, 2) identifying genes within taxa that correspond to these metabolic pathways, and 3) confirming pathway activation through transcriptomic data. Cases in which interacting genes, transcripts, and metabolites covary (or are co-absent) provide “guilt-by-association” evidence of new functional annotations, novel biochemical pathways, or metabolite contributions from non-microbial sources.
To determine which microbes are active, we will use a combination of ‘omics approaches on newly collected samples that include metagenomics, metatranscriptomics, and metabolomics. From this, we can identify live bacteria (DNA) based on those that are transcriptionally active (RNA). We can further determine which metabolites are being produced by 1) associating taxonomic profiles with metabolites, 2) identifying genes within taxa that correspond to these metabolic pathways, and 3) confirming pathway activation through transcriptomic data. Cases in which interacting genes, transcripts, and metabolites covary (or are co-absent) provide “guilt-by-association” evidence of new functional annotations, novel biochemical pathways, or metabolite contributions from non-microbial sources.
To construct microbial signatures for different built environments, we will use the hitting set algorithm generated by Eric Franzosa, a research associate in the lab. This was developed to identify human hosts by their persistent microbial strain combinations, and in this study, we will test how these “codes” may identify individual built-environments as well. In brief, we will determine sets of features (taxa, genes, functions) that are unique to train lines and stations. We can then evaluate how stable these microbial codes are between our previous and new collections. Together, these features may provide a strategy for characterizing recently-visited environments using microbes newly acquired by an individual.
Generally, it is believed that microbes in the built environment originate from their human inhabitants and surrounding environments. This was the case in our last study, e.g. contrasting the presence of environmental taxa on indoor versus outdoor touchscreens, the prevalence of skin microbes on trains, and the presence of vaginal microbes on seats but not seatbacks. In terms of macroecology, urbanization frequently reduces biodiversity,1 though there are select animals such as pigeons and sparrows that thrive in (or more typically near) the built environment.2 Analogously, there may also be specific microbes that thrive and contribute to the built environment, which will be increasingly important to understand as more people move to cities. Overall, this study will help identify these microbes and the mechanisms they use to survive, as well as determine how stably they inhabit built environments across time.
Footnotes
1. Sol, D., Lapiedra, O., Gonzalez-Lagos, C. (2013). Behavioural adjustments for a life in the city. Animal Behaviour, 85, 1101-1112. http://dx.doi.org/10.1016/j.anbehav.2013.01.023
2. Kark, S., Iwaniuk, A., Schalimtzek, A. and Banker, E. (2007), Living in the city: can anyone become an ‘urban exploiter’?. Journal of Biogeography, 34: 638–651. doi:10.1111/j.1365-2699.2006.01638.x