(this post was written by group #4 as a writing assignment)
With a minor setback from last week’s inability to efficiently wash and quantify the amplified DNA, we came back this week with a stronger mindset to pick ourselves back up and turn our failures into a positive. At the start of this week’s class, we miraculously had quantifiable DNA to send off to the Sanger sequencing facility, and get forward and reverse reads of our bacterial DNA . (Disclaimer: a little magic may or may not have happened behind the scenes by our professors) As a result, during this week’s class our goal was to make a consensus sequence from the Sanger sequencing results and BLAST it to finally hypothesize what species we could possibly be dealing with.
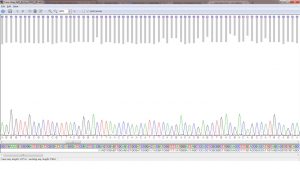
With that in mind, we stepped into the computer lab and started the next part of finding the identity of the bacterial DNA that we had isolated from the abalones. The first thing we did was use SeqTrace, a program that allowed data to be open sourced, free, as well as current with the technology today. Through SeqTrace, we were able to match the forward and reverse sequences of our results from the Sanger Sequencing, cut off the crappy end parts, and get a consensus of the most probable sequence that we isolated. We then uploaded these results to a Standard Nucleotide Blast at NCBI to match to our DNA to known bacterial species. All of us got a match, picked our top 3 results based on the closeness of nucleotide bases, and saved them for next week’s class, where we will use that information to create phylogenetic trees and hopefully be one step closer to figuring out our bacterial species and possibly provide an answer to the abalone withering syndrome.
An important lesson we learned this week is that despite science always seeming to follow a set plan and expecting outcomes to go smoothly, we found that experiments do not always go as planned and that errors do happen. Something we often forget is that experiments are not always error-prone, they are called experiments for a reason. But the important thing in science is that we learn to come back with a stronger mindset, correct our problems, and get back to doing what we love, testing and expanding the boundaries of our world today.
So with that, we have one question that we have for experienced scientists: Do errors or setbacks happen in your lab/projects and do you have any advice about persevering through the struggles? Feel free to post your stories below, we’d love to hear them!!! or Does extreme DNA contamination happen often and does this ever completely ruin your data?