Guest post from Trent Northen of LBNL. I wrote to Trent recently when I had seen a paper of his that I found fascinating: Exometabolite niche partitioning among sympatric soil bacteria. So I asked if he would be willing to write a guest post about it. And so – he did. It seems like the issues discussed in this paper could apply to many microbial communities and thus might be of general interest. Anyway – here is the blog post below:
The Northenlab (northenlab.org; @northenlab) has been profiling metabolites from microbes and microbial communities since 2008. This never gets old. Several years ago we started looking at uptake and release of metabolites in addition to profiling intracellular metabolites. We wanted a good model system to apply these methods to and learned about biological soil crusts (biocrusts) from Ferran Garcia-Pichel at ASU, and started a collaboration with his group, Eoin Brodie, Cheryl Kerfeld and Aindrila Mukhopadhyay at Berkeley Lab.
These biocrusts are top-soil microbial communities and are typically found in plant interspaces. They roughly cover 20% of continental areas and were most likely even more prominent before the explosion of land plants in the Devonian. It is accepted that members of the filamentous Microcoleus sp. genus of cyanobacteria, are the pioneer organisms that bind the sand together, stabilizing the soil from erosion enabling succession into fairly diverse communities (some ~2000 Operational Taxonomic Units (OTUs)).
The competitive exclusion principle states that pure competition cannot exist, so no two microbes can occupy the same niche. Thinking of the 2000 OTUs in the Biocrust, it is hard to imagine how there are enough niches in ~1mm thick community growing in a simple sand matrix. In our recent paper (http://bit.ly/1QwzPMS) we describe how when we analyzed both the spent media from Microcoleus vaginatus and biocrust pore water we find hundreds of metabolites and wondered if microbial feeding preferences for these exometabolites could contribute to niche partitioning.
Biocrust Exometabolite Niche partitioning? Do microbes from biocrusts pick their favorite food from the biocrust metabolites buffet?
To investigate this we selected M. vaginatus and 6 phylogenetically diverse heterotrophs and sympatric isolated by the Brodie Lab. The isolates were grown in rich media made by pooling the extracts of the isolates (soil DOM is thought to be largely composed of microbial products) and used LC/MS based exometabolomics to compare the inoculated vs. control media to determine what each isolate consumed from the media. We were surprised to find that M. vaginatus was very leaky compared to other Cyanobacteria, in fact most of its intracellular metabolites were detected in the spent minimal media (though many abundant metabolites were not detected suggesting that this was not from cell lysis). Since this primary producer does not fix nitrogen, these ‘exudates’ may be important in stimulating heterotrophic nitrogen fixers.
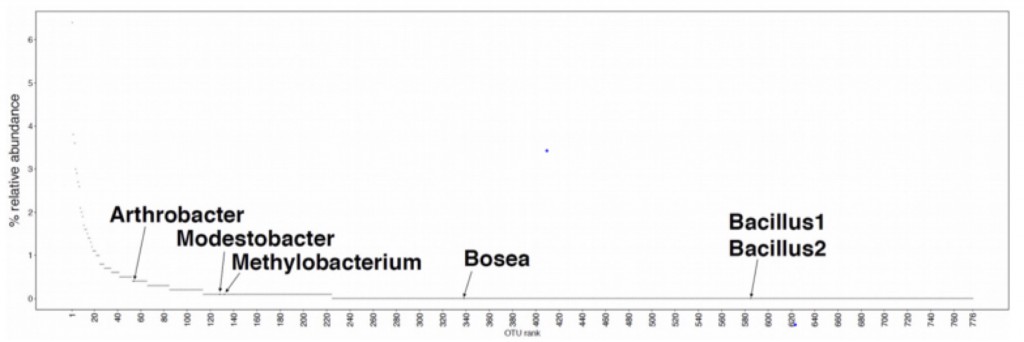
We expected that most if not all of the bacteria (esp. the two Bacilli) would be generalists and would consume essentially everything. However, this is not what we found. Only two of the 470 metabolites characterized (glutamate and an unknown metabolite in positive mode) were consumed by all 7 isolates (including M. vaginatus), only 7 by all of the heterotrophs and about 40% weren’t consumed by any of the isolates. Interestingly M. vaginatus consumed the widest range of (its own) metabolites.
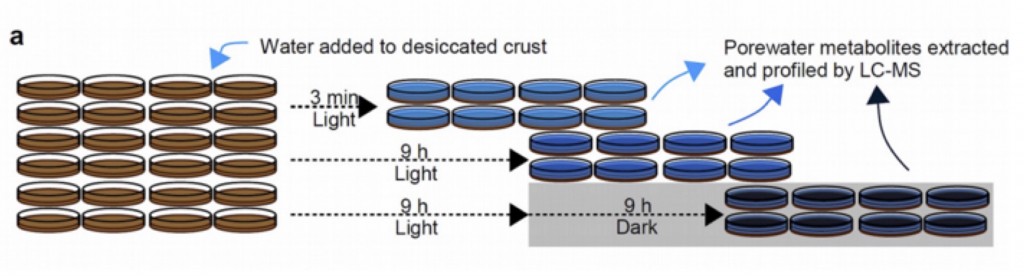
These results were compared with time-series profiling of desiccated intact biocrusts. Here the crusts were placed in petri dishes and saturated with water in the lab under a light/dark cycle. Soil water was collected at 3min, 9 h and 18 h following wetting and analyzed using LC/MS metabolomics. Most of the metabolites detected in the spent complex media of the pooled isolates was also detected in the biocrust pore water. Many metabolites consumed by the isolates were also consumed in the intact crusts. However, there were numerous examples of metabolites consumed by the biocrust but not by the isolates.
This result and a similar study on subsurface microbes suggest that exometabolites may contribute to diversity by minimizing competition. That is microbes could occupy the same physical niche but use different substrates, pick different food from the metabolites buffet. Though many more isolates and environments need to be considered before generalizing this finding. We also detected many metabolites that were depleted by one and released by another, suggesting possible cross-feeding and this could potentially help explain the small degree of substrate utilization overlap.
Last month we published another paper on these biocrusts. Here, we studied another group of cyanobacteria in the biocrusts producing ‘sunscreens’ that darken the soil, decreasing its albedo (ability to reflect light and energy) and actually cause an increase in the land-surface temperature (http://bit.ly/1Pe7g5K). This work was led by Estelle Couradeau in Ferran’s lab. This work built on earlier work by Ferran’s group (http://bit.ly/1QwHmv6) showing that at a continental scale biogeographic study of cyanobacteria across the western US, that temperature largely determined which species of Microcoleus sp. dominated. In hot deserts, Microcoleus steenstrupii dominated and in colder deserts Microcoleus vaginatus was dominant. His group isolated many strains from each species and found that indeed the M. steenstrupii grew better at temperatures about 10 °C higher than and M. vaginatus.
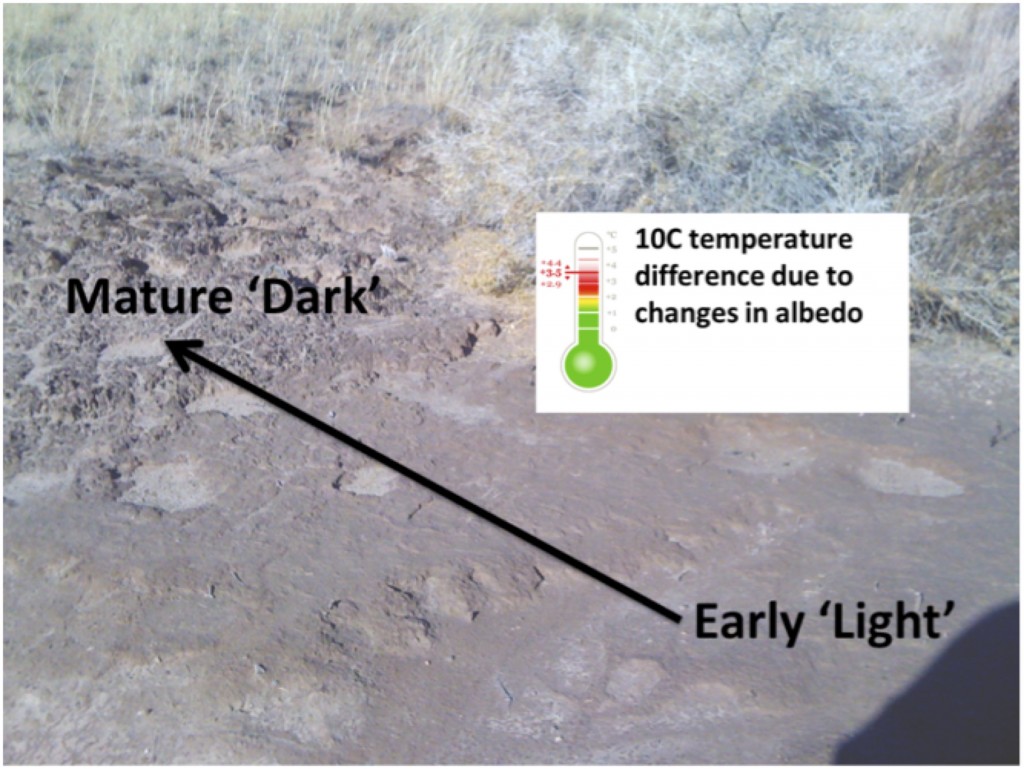

Interestingly near Moab UT these two species co-exist and typically M. vaginatus is dominant in the early successional ‘light’ crusts and M. steenstrupii in the more mature ‘dark’ crusts. We wondered if the dark color might result in higher soil temperatures that could select for the hot adapted M. steenstrupii and Estelle characterized the surface temperature (under full-sun) across a maturity transect (space-for-time) going from very ‘light’ early stage crusts to mature ‘dark’ crusts and found that the temperature increase 10 °C! In collaboration with Ulas Karaoz and Ulisses Nunes da Rocha from Eoin Brodie’s lab and others they sequenced a transect of crust maturity and found that indeed in this small area the community does shift from M. vaginatus to M. steenstrupii dominated, just as observed on the continental scale in the earlier biogeographic study.
Estelle extracted the pigment from these Biocursts and found that this dark color is a result of the accumulation of the ‘sunscreen’ pigment scytonemin (first described by Ferran when he was at the University of Oregon produced by nitrogen-fixing cyanobacteria of the order Nostocales, both the abundance of these organisms and the pigment itself was found to increase with maturity. Thus we find a fractal type behavior where we see patterns of small scale temperature gradients driving shifts in primary producers nested within larger latitudinal scale shifts. And more importantly this contribution highlights for the first time the role microbial communities as important players of soil albedo and surface temperature, a role that was previously only credited to land plants.
Some responses from Twitter: