I’m working on a manuscript where we analyze OTU co-occurrence patterns using network analyses as well as the traditional approaches (ordination, alpha diversity, etc.). This means that we have to include some equations in our methods and so for the first time, I am using Authorea and LaTeX. I am getting used to working with the …
Here are my notes from day 3 of the Mothur Workshop that is taught by Pat Schloss (pdschloss at gmail.com) at a hotel that is conveniently located near the Detroit airport. Of all of the bioinformatic workshops I have attended, this group of students was my favorite so far. We had a lot of fun …
Here are my notes from day 2 of the Mothur workshop taught by Pat Schloss (pdschloss at gmail.com) in December 2015. For those who are interested in learning to use Mothur for microbiome studies, Pat will be teaching another one in February. Mothur is better for bacterial characterization than eukaryotes because the sequences are aligned before OTU clustering. This …
Pat Schloss (pdschloss at gmail.com) offers excellent workshops on the Mothur software for analyzing 16S rRNA data for bacteria. He’s just announced the next one (February 8 to 10, 2016 near the Detroit airport). I had the pleasure of attending the December workshop. A diverse and international group attended the workshop, with many folks who are interested in the human microbiome. …
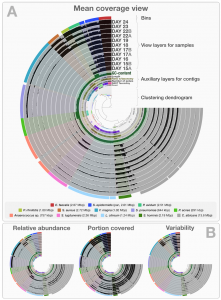
Blog post prepared jointly by Andrew Doxey (@acdoxey) and Josh Neufeld (@joshdneufeld) The “aquariome” Back in 2013, as part of a project assessing aquarium microbial communities and their role in nutrient cycling, Laura Sauder (graduate student in the Neufeld lab) sequenced a shotgun metagenomic library from a freshwater aquarium biofilter that was installed on this …
Three weeks ago I stood in front of the 60 attendees of the STAMPS course and asked, “How many of you are currently working with shotgun metagenomes?” Ten to fifteen people raised their hands. In contrast, almost all had their hands in the air when I asked how many were expecting to work with shotgun …
Dear metagenome method developers, The first challenge of the Initiative for the Critical Assessment of Metagenome Interpretation (CAMI) begins right now! Over the last three months, we received valuable feedback from the community playing with our toy data sets. We incorporated many of your suggestions, thanks again! Today, we proudly release the official data sets …
Aaron Darling, faculty member of University of Technology Sydney (who used to work with me here at UC Davis) has a very important and interesting read on his blog: Not so fast, FastTree. In it he discusses some informatics archaeology he did (digging around in some code) regading the program FastTree which many researchers have been using …
SciPy 2015 (Scientific Computing with Python) is coming up in Austin, TX this July 6-12. I attended SciPy last year for the first time to present on scikit-bio (see my talk here), and thought it was an excellent meeting. It was great to spend a week talking about software and software development, which isn’t the …
Compiling some of the more interesting tools I have seen recently. Some I have plyed with but most I have just looked at the papers briefly. Microbiome | Abstract | VizBin – an application for reference-independent visualization and human-augmented binning of metagenomic data. Global biogeographic sampling of bacterial secondary metabolism GrammR: Graphical Representation and Modeling …