(This is a guest blog post, written by Madhan Tirumalai and George Fox, authors of the publication)

Madhan R Tirumalai and George E. Fox
Department of Biology and Biochemistry, University of Houston, Houston, USA.
The Spacecraft assembly facilities are a special kind of the built environment. Their uniqueness stems from the fact that space agencies have to abide by the provisions of the 1967 Outer Space Treaty as well as current planetary protection policy, which depending on the mission stipulate strict cleaning protocols in order to avoid or minimize forward contamination. In order to achieve extremely low levels of particulate, high efficiency filtration technology is used in combination with controlled moisture & temperature conditions. Surfaces in the clean room are decontaminated using a variety of techniques including desiccation, hydrogen peroxide treatment, and UV radiation. The result is a special ‘extreme’ habitat that is perfect for natural selection for multi-resistant bacteria. Even as far back as 1967, microbial assay of the Explorer XXXIII Spacecraft (Anchored Interplanetary Monitoring Platform, or AIMP) showed substantial viable microbiological burden. Thus microbial contamination in such environments is unavoidable.
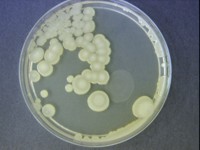
In recent years, a number of novel bacterial species that exhibit elevated resistances to desiccation, peroxide, UV, and gamma radiation have been isolated. Endospore forming Bacillus species feature prominently amongst this inventory of spacecraft assembly cleanroom microbiome. Endospores can remain dormant for years until they germinate to grow vegetatively upon reactivation. With modern sequencing technology, the genome sequences of these uniquely radiation resistant spore producers, can be studied and compared with those of very closely related non-resistant spore producers. Such comparisons help yield differences that could help unravel the spore resistance puzzle.
Two endospore producers, namely, B. safensis FO-36bT(isolated from clean-room air particulate, JPL-SAF, 1999) and, B. pumilus SAFR-032 (isolated clean-room airlock, JPL-SAF, 2001), produce spores that are extremely UV (254nm UV) resistant and peroxide resistant. The genomes of these two strains were sequenced and compared with that of B. safensis JPL-MERTA-8-2 (isolated from the Mars Odyssey Spacecraft and associated facilities at the JPL and later also found on the Mars Explorer Rover (MER) before its launch in 2004) and that of the non-resistant spore-producing B. pumilus ATCC7061T. Technically, these four strains are ‘microbial’ close cousins, of which, two produce resistant spores (FO-36bT and SAFR-032), while one (JPL-MERTA-8-2) grows well in the International Space Station, but its resistance properties are not known, and the remaining ‘cousin’ ATCC7061T is not resistant!
To put this in rudimentary terms, comparing these genomes is akin to hiking the landscape of say, the Rocky Mountains and mapping the contours of the similar looking mountains. The difference in the genome comparison lies in that we are looking at genome architecture, the presence or absence of genes, shared genes, genes unique to a certain strain and sequence level differences in specific gene families amongst the genomes, all of which in such comparative studies gives an insight into the differences and how they could be linked to the phenotype.
During this process, four ways of comparisons were made, viz., gene vs gene, protein vs protein, gene vs genome, and, protein vs proteome between not only the four genomes mentioned here, but also including 61 other genomes of strains identified and named either B. pumilus or B. safensis in the public database (NCBI).
By doing so, we identified 9 genes that are totally unique to B. safensis FO-36bT, and examining the location of these 9 genes, revealed that four of them are part of a phage insertion element, namely, the Bacillus bacteriophage SPP1. Thus, we looked for similar phage insertions in the other 3 as well as 61 other genomes. The section of the insertion containing the four genes unique to B. safensis FO-36bT are absent in all the other genomes.
Phages play major roles in the evolution of bacteria by contributing to the genetic variability between closely related bacterial strains. Such variability is often implicated in the phenotypical differences such as bacterial pathogenesis. Bacteriophage-mediated horizontal gene transfer enhances bacterial adaptive responses to environmental changes. Furthermore, phages shunt genes that could directly impact the phenotype between related strains or between phylogenetically distant strains via horizontal gene transfer (HGT). All of these evolutionary events have implications for selection and fitness.
The SPP1 phage (harboring the four unique genes of B. safensis FO-36bT) is well-known for its ability to mediate transfer of any gene from one bacterium to another. This raises the following questions: How are these genes linked with the phenotype of B. safensis FO-36bT? And what are the possibilities of them getting shunted around to other non-resistant strains?
Previous genome comparisons of non-sporulating radiation resistant bacteria yielded significant insight to the origins of resistance. For example, in the case of the super radiation resistant organism Thermococcus gammatolerans, the high radio-resistance has been linked to proteins that remain to be characterized. Given that the 10 genes are unique to the B. safensis FO-36bT, these are potential candidates that could be investigated further for their roles in the strain’s resistance properties.
Understanding the complex dynamics of microbial ecology in any environment, would require a combined systems approach including, comparative genomics, transcriptomics, and proteomics. This paper talks about a fundamental comparative genomics approach towards identifying potential candidate genes with possible roles in the radiation spore resistance properties of a Bacillus strain from the spacecraft assembly environment.
The bigger question is whether these spores may ride piggyback on spacecraft. As of now, even if they were to indeed hitch a ride, they may not possess the ability to survive under extraterrestrial conditions such as those found on Mars. However, it is a matter of concern that there always seem to be a background level of such spore producing strains and making an inventory of the same is vital, since getting rid of 100% of them is unrealistic. Such an inventory would help in fine-tuning what NOT to look for in samples retrieved from elsewhere.
The genomes of these organisms on the other hand are treasure troves to examine and analyze and the knowledge acquired from this process can further be applied towards understanding the genome complexity of strains from other such confined habitats.
References:
- Tirumalai MR, Stepanov VG, Wünsche A, Montazari S, Gonzalez RO, Venkateswaran K, Fox GE: Bacillus safensis FO-36b and Bacillus pumilus SAFR-032: a whole genome comparison of two spacecraft assembly facility isolates. BMC Microbiology 2018, 18(1):57.
- Rummel JD: Planetary protection policy overview and application to future missions. Adv Space Res 1989, 9: 181–184.
- Powers EM: Microbiological burden on the surfaces of Explorer 33 spacecraft. Appl Microbiol 1967, 15(5):1045-1048.
- Tirumalai MR, Rastogi R, Zamani N, O’Bryant Williams E, Allen S, Diouf F, Kwende S, Weinstock GM, Venkateswaran KJ, Fox GE: Candidate genes that may be responsible for the unusual resistances exhibited by Bacillus pumilus SAFR-032 spores. PLoS One 2013, 8(6):e66012.
- Coil DA, Neches RY, Lang JM, Brown WE, Severance M, Cavalier D, Eisen JA: Growth of 48 built environment bacterial isolates on board the International Space Station (ISS). PeerJ 2016, 4:e1842.
- Cano RJ, Borucki MK: Revival and identification of bacterial spores in 25- to 40-million-year-old Dominican amber. Science 1995, 268(5213):1060-1064.
- Yasbin RE, Young FE: Transduction in Bacillus-subtilis by Bacteriophage Spp1. J Virol 1974, 14(6):1343-1348.
- Zivanovic Y, Armengaud J, Lagorce A, Leplat C, Guerin P, Dutertre M, Anthouard V, Forterre P, Wincker P, Confalonieri F: Genome analysis and genome-wide proteomics of Thermococcus gammatolerans, the most radioresistant organism known amongst the Archaea. Genome Biol 2009, 10(6):R70.
