Just a quick post here. There is a paper of interest I thought I would call attention to:
Source: SILVA, RDP, Greengenes, NCBI and OTT — how do these taxonomies compare? | BMC Genomics
by Monika Balvočiūtė and Daniel H. Huson
Abstract:
Background
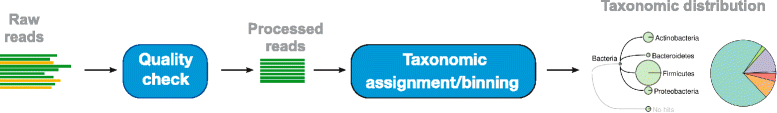
A key step in microbiome sequencing analysis is read assignment to taxonomic units. This is often performed using one of four taxonomic classifications, namely SILVA, RDP, Greengenes or NCBI. It is unclear how similar these are and how to compare analysis results that are based on different taxonomies.Results
We provide a method and software for mapping taxonomic entities from one taxonomy onto another. We use it to compare the four taxonomies and the Open Tree of life Taxonomy (OTT).Conclusions
While we find that SILVA, RDP and Greengenes map well into NCBI, and all four map well into the OTT, mapping the two larger taxonomies on to the smaller ones is problematic.
See the paper for more details. Their software for mapping taxonomies is available here: http://ab.inf.uni-tuebingen.de/software/crossclassify/