Infants born via c-section have a microbiome community composed mostly of skin bacteria [1-3], but the source of these skin bacteria is unknown. People quickly shed bacteria into their environment, leaving their own bacterial signature in a room within hours [4]. Do hospital operating rooms harbor skin bacteria that could colonize c-section delivered infants? A study published earlier this week by Shin et al. in Microbiome addresses this question.
To characterize the microbiome of operating rooms, four hospital operating rooms in two different cities, New York, New York and San Juan, Puerto Rico, were sampled immediately after c-section births using culture-dependent and independent techniques. Dust was also examined under the microscope for skin cells. Sampling sites included walls, floors, return air vents, counters, cribs, and lamps above the operating table. Infant skin was sampled 1-7 days after birth in 3 locations and compared to operating room samples as well as human oral, fecal, and vaginal microbiome samples from the Human Microbiome Project datasets.
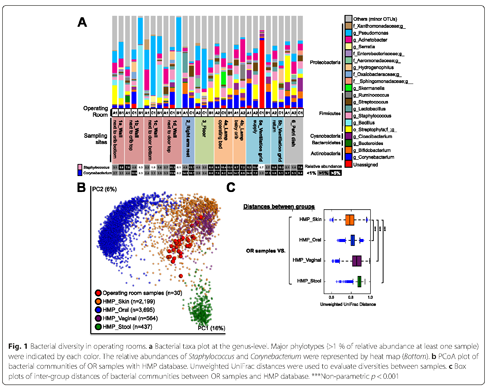
Operating room samples were most similar to human skin bacteria, as compared to oral, fecal, and vaginal microbiome samples (Figure 1). All hospital samples were dominated by Staphylococcus and Corynebacterium, common human skin bacteria. Lamps had the highest abundance of these two skin bacteria, including live cells that could potentially be transmitted to infants if the lamp is moved. Microscopy revealed human skin flakes in operating room dust samples. Bacterial beta diversity showed microbiome communities clustering by hospital and even differences between two different operating rooms in one hospital. Alpha diversity did not vary between hospitals. These results suggest that there may be differences due to either occupants in an operating room or cleaning regimes.
SourceTracker [5] results suggested that the skin microbiome of c-section delivered infants has bacteria obtained from the operating room. In contrast, the microbiome composition of vaginally-born infants is more similar to that of their mother’s vagina and gut microbiome. C-section infants are more likely to maintain a different microbiome than vaginally born infants throughout the first year [2, 3, 6] and perhaps even into adult hood [7]. Since C-section infants are also more likely to develop allergies, asthma, and digestive system disorders, understanding the source of the bacterial inoculum for C-section babies is important to determine. This paper is a first step towards understanding the potential role of the operating room in the colonization of infants during c-section delivery.
REFERENCES
- Dominguez-Bello MG, Costello EK, Contreras M, Magris M, Hidalgo G, Fierer N, Knight R: Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proceedings of the National Academy of Sciences 2010, 107(26):11971-11975.
- Bäckhed F, Roswall J, Peng Y, Feng Q, Jia H, Kovatcheva-Datchary P, Li Y, Xia Y, Xie H, Zhong H et al: Dynamics and Stabilization of the Human Gut Microbiome during the First Year of Life. Cell Host & Microbe 2015, 17(5):690-703.
- van Nimwegen FA, Penders J, Stobberingh EE, Postma DS, Koppelman GH, Kerkhof M, Reijmerink NE, Dompeling E, van den Brandt PA, Ferreira I et al: Mode and place of delivery, gastrointestinal microbiota, and their influence on asthma and atopy. J Allergy Clin Immunol 2011, 128(5):948-955 e941-943.
- Lax S, Smith DP, Hampton-Marcell J, Owens SM, Handley KM, Scott NM, Gibbons SM, Larsen P, Shogan BD, Weiss S et al: Longitudinal analysis of microbial interaction between humans and the indoor environment. Science 2014, 345(6200):1048-1052.
- Knights D, Kuczynski J, Charlson ES, Zaneveld J, Mozer MC, Collman RG, Bushman FD, Knight R, Kelley ST: Bayesian community-wide culture-independent microbial source tracking. Nat Meth 2011, 8(9):761-763.
- Arrieta M-C, Stiemsma L, Amenyogbe N, Brown E, Finlay B: The Intestinal Microbiome in Early Life: Health and Disease. Frontiers in Immunology 2014, 5.
- Goedert JJ, Hua X, Yu G, Shi J: Diversity and Composition of the Adult Fecal Microbiome Associated with History of Cesarean Birth or Appendectomy: Analysis of the American Gut Project. EBioMedicine 2014, 1(2—3):167-172.
I looked for but could not find the building science in this article. Can you help me? Thanks.
sorry – I’ll remove the tag.
Anne – not sure if Hal was being sarcastic or snippy there or trying to be helpful. Regardless, I am glad you used a tag. Tags are very helpful for posts for many reasons (for searches, for sorting, for people to find similar posts and so on). I note – of course – there is no way to have tags and categories cover everything so sometimes it is necessary to use tags that don’t exactly match what is in the post just like you did.
I have taken the liberty of adding a bunch of tags to this post as well as created a new category “Built Environment” with a sub category “Buildings” so that we can categorize posts like this which are clearly about the built environment.
Thanks again for the post.