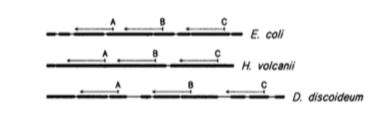
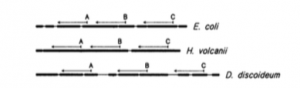
I got this email a few days ago from Jed Fuhrman. He had sent it to a group of people working on mcirobial diversity and he encouraged people to share it. I asked and he approved posting it here and I thought it would be of interest. (The image above is from Lane et al. 1985 -see below).
Hi everyone, (let me apologize in advance if you think this is spam, sent mostly to people associated with EMP, OSD, JGI, and TARA)
Our paper, Parada et al., describing the mock-community and field sample evaluation of the original “EMP” and alternative (515-926) ssu rRNA primers for marine microbiome studies, is now out in Environmental Microbiology, advanced online publication. The link is
http://onlinelibrary.wiley.com/doi/10.1111/1462-2920.13023/abstract?campaign=wolacceptedarticle
Main points:
- The original EMP primers(515-806) also used by OSD, JGI, and many others greatly underestimate the highly abundant marine SAR11 cluster in particular (plus other groups) and overestimate Gamma Proteobacteria in particular (plus other groups) in marine samples. When we use a mock community of 27 marine clones (uneven rank-abundance distribution crudely simulating marine plankton) the r^2 of “observed vs expected” with these primers is only ~0.5 .
- The bias against SAR11 is about 2- to 5-fold worse in field samples than in the relatively “simple” mock communities (for reasons we can only speculate —maybe primer competition on very diverse samples?), suggesting the real r^2 in the field is probably worse than 0.5 with marine plankton. Note that the EMP primers have a known mismatch to most SAR11, but we found that even sequences perfectly matching the primer could show strong biases.
- Our recommended alternative primerset (E. coli numbers 515Y-926) has an r^2 of observed vs expected of 0.95 with the same 27-member mock community, so it has far fewer biases. It also encompasses the entire original EMP-primer amplicon, allowing direct comparisons of the results.
- The alternative primer set PCR products include an additional variable region, so the results have usably higher phylogenetic resolution, with ecological significance. We show examples where natural bacterial sequences from our San Pedro Ocean Time-series include taxa that are distinguished only with the longer 515-926 product (i.e. the same over the 515-806 region, different between 806 and 926) and these can have very different patterns of variation over time.
- The alternative primer set also hits eukaryotic 18S rRNA well.Initially we were concerned that eukaryotic sequences would dominate (swamp) some samples, but even in >1 um marine plankton samples from a spring bloom (DNA dominated by eukaryotic phytoplankton, plus other protists and some metazoa) average only 17% eukaryote 18S sequences (highest was about 1/3 18S). Note that in phytoplankton-dominated samples, a large fraction of all 16S sequences come from chloroplasts anyway. The eukaryote 18S amplicons are longer, so forward and reverse reads do not overlap even in typical 2 x 300 MiSeq outputs. Therefore these sequences are lost from informatic pipelines that require overlap. However, including them opens up the possibility of a good coverage (>500 bp, phylogenetically informative) of eukaryote 18S sequences in the same run as 16S — allowing for a true 3-domain total microbiome coverage all at once. Note that if desired, the longer 18S amplicon could be separated, pre-sequencing, from the 16S, e.g. by a gel or high-throughput bead separation. Yes, we are aware that the unexpectedly low 18S percentage in eukaryote-dominated samples suggests a strong bias against these sequences, probably because of the longer product. It benefits people interested mostly in bacteria and archaea. We have not yet evaluated the extent of the biases with mixed 16S and 18S mock communities.
- We corrected a mismatch to the Thaumarchaea in the original EMP 515 primer, and found that even in deep sea samples with many Thaumarchaea, it hardly changed the proportion of sequences from those taxa (slight increase when 806 reverse is used, no significant change when 926R is used); however the 515-926 primers yielded many more Thaumarchaeal sequences than the 515-806 primers.
- The upshot is that in-silico evaluation of primers is necessary but not sufficientto determine their suitability — complex mock communities and careful examination of field samples is also needed. Biases vary considerably depending on the composition of the samples, so they DO NOT NECESSARILY “CANCEL OUT” when comparing samples, even though they are reproducible in replicate samples. As microbiome studies become more and more routine, it becomes more important to avoid significant biases whenever possible, in large part because the smaller the biases, the less we have to worry about to what extent they are the same from sample to sample. Yes, there are many other biases besides primer-based ones, in sampling, extraction, copy number, etc. But with applications like PICRUSt being used to predict the total metagenome from the ssu rRNA, it behooves us to provide an accurate-as-possible picture of the microbial communities, and avoid known biases. We know the alternative primers have mismatches to many environmental sequences, and welcome suggestions for other alternatives, evaluated in similar ways. We study marine microbes and we evaluated these primers specifically for them, picking primers that hit as many known marine taxa as possible. Interesting note: the 515-926 primers also correspond (with a few minor changes) to two of the three original “universal” ssu rRNA reverse-transcription primers originally published in 1985 before the advent of PCR, by Lane et al. (Pace lab, of course) in PNAS. Their third primer was 1392R, also still an excellent choice, probably soon suitable with the 515F as sequencing platforms yield even longer products.
- We did this work because as we (slowly) transitioned to NGS 16S sequence-based studies, we were concerned that few labs had systematically evaluated the methods, with “standards,” blanks, etc. (although the well-funded HMP did this, somewhat differently, for human microbiomes some years ago). My former grad student (now postdoc) David Needham deserves particular credit for pushing this and creating the mock communities, and my student Alma Parada did the bulk of the work for this paper. We originally thought it would be an easy task, but the unexpected variations in biases (among other issues) made it tricky.
- This work is not meant to be a general criticism or indictment of any other studies done in the past, but rather a forward-looking recommendation.
I encourage you to forward this message to anyone you feel may benefit from it.
Best,
Jed
Norm Pace then wrote a response in regard to general and historical information on rRNA primers. His main point was that Lane et al. 1985 included a discussion of the “927” primer and that Jed had not discussed this in the current work.
And Jed agreed with Norm’s points:
Hi Norm,
You are completely right that Lane et al. (1985) nailed it on the best locations for universal primers, and we cited you when we used them starting in our 1992 Nature paper. The current primers vary somewhat: 3 bases shorter than Lane et al.’s equivalent to the 515F, and there are two extra ambiguities in the 906 primer. In the current work we cited papers reporting the exact versions we actually used, but you are right that you and your group deserve credit for the incredible foresight you had. I’ll see if we can add the PNAS citation in the proofs.
Best,
Jed
Jed, there are many of us working with microbial diversity that cannot follow up with the progress of molecular methods in biodiversity studies. Could you please explain your contribution so we can understand the consequences of primer biases on revealing microbial diversity? Does your study contribute to sort artefacts/eukaryotes out of the marine “rare biosphere”, or the “grey matter” bulk? Does it show that we are overestimating certain taxa, and underestimating others?