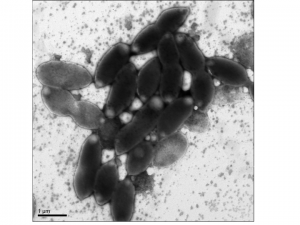
After way more work than I imagined when we embarked on this project, we are almost ready to submit our first description of a new bacterial species, Kirrobacter mercurialis. Not exactly earth-shattering, given that about 40 bacterial species descriptions come out every month in the International Journal of Systematic and Evolutionary Microbiology (IJSEM). However, our lab tends to focus on the grand-scale of biology (metagenomics, 16S surveys, population biology) or on bioinformatics (phylogenomics, genome assembly, software development). We’re not really a “microbiology” lab in any sort of classical sense. In fact when I joined the lab I had to order petri dishes and microscope slides since the only wet lab work was DNA extractions and library preparations. However, I always wanted to describe a new species and the opportunity arose through Project MERCCURI (a.k.a space microbes) where one of the isolates we grew up only showed 95% 16S identity to any cultured bacterial species. With the help of a series of undergraduates in the lab (Jennifer Flanagan, Andrew Stump, and Alex Alexiev) we began to try to figure out what this process would entail.
I started off by reading a number of existing species descriptions, and in my naivety was struck by several things:
DNA-DNA hybridization is still considered a gold standard to measure the relatedness of a new species to close relatives. IMHO this is insane, given the fact that this is a technically difficult, expensive (you usually have to buy DNA for comparison from culture collections) technique that is, at best, a fairly gross measure of relatedness. In our case the very first thing we did was to sequence the genome… we could compare that to other genomes at a fraction of the cost and effort. Fortunately, our bug was distant enough from close relatives that it would have been pointless to even try the DNA-DNA hybridization.
Speaking of having the genome, there are a number of aspects to classical microbiology that can benefit enormously from having this kind of data. For example, members of the family where Kirrobacter is located contain carotenoids, so measuring these is a standard part of a species description. This process entails a methanol extraction of cell lysate, followed by analysis of absorption peaks on a spectrophotometer. Or in our case, I spent 30 seconds looking up the carotenoids in the RAST annotation of the genome. Done. Another interesting case for us was with flagella. We performed motility tests (negative) and examined the bug using both light and electron microscopy during both exponential and stationary phase. Never once did we observe flagella, yet the genome contains 30 flagella genes (compared to 0 for another non-motile close relative). This suggests that the picture is more complicated than just saying “no flagella”.
Just to be clear, I think there is very significant value in doing species-level classical microbiology! That’s partly why I started this… I think I looked at one too many lists of taxa in QIIME and wondered “but what are they *doing*?”. There are many aspects to the biology of a species that can’t (yet) be determined from the sequence. For example; morphology, temperature range, salt tolerance, etc. Many of these are important in understanding what niche a species might occupy, or what it might be actually doing in a community. I think far more work on cultured microbes is needed before we can start to paint complete pictures of complex microbial communities in the real world.
That being said, I don’t get the impression that that the requirements to describe a new species reflect what’s really important, given access to modern tools like cheap genome sequencing and good sequence-based taxonomy. For example, all bacterial species descriptions devote a lot of space and either pain/$$$ to characterizing the fatty acids in the membrane. Historically, this was an important part of determining taxonomy. Nowadays there are important arguments between various schools of thought as to whether a determinative taxonomy should come from 16S, a collection of housekeeping genes, or whole genomes… but no one (that I’m aware of) is discussing fatty acid profiles. And functionally, I’m not sure there’s much value in knowing that Kirrobacter mercurialis has a normal amount of C18:1ω7cis compared to other members of the family, but has more C16:0.
For this, my first involvement in a species description, I wanted to do it “right” and use the appropriate classical tools. The only exceptions to this were the DNA-DNA hybridization as mentioned above and publishing in the closed-access IJSEM (fortunately you can petition for the name to be mentioned in IJSEM, which still qualifies for official standing in nomenclature). I’d like to do another species description sometime that focuses on information from the genome, and those characteristics likely to be most useful to someone who either wanted to culture the organisms or understand something about it’s biology. Sadly, lacking the classical measures would probably mean that the species would never have official standing in nomenclature.
I absolutely agree with you on the DNA-DNA hybridization rule. I did a formal species description for Nautilia profundicola back in 2008. We also had the whole genome but were turned down in our request to do an in silico comparison to nearest neighbors. We ended up paying DSMZ to grown up the two strains and do the DNA-DNA hybridization. Seems a pity it is still this way.
Dear Jonathan
Nice to have your opinion on what is important for a new taxa description. I was happy to see that you as one of the highly regarded molecular experts mentioned physiological characterization as important for taxa description, such as temperature profiles, pH-profiles, growth rates/doubling timesetc. I am fighting–and got in trouble for that– that all the biochemical characterization required by some of the IJSEM power people such on e.g., phospholipids etc. are nice if you can have them, but should not be an absolute requirement. Such assays –in many cases depending on the labs possibilities– has to be done by DSMZ for lots of $$. But that the same persons do not pay any of the physiological characterization. Myself, like many other microbiologists are more interested in what the new taxa can do and how to grow it optimally. The argument that those data depend so much on growth conditions and are to variable is in my opinion not really valid because the same is true e.g., for the lipid profiles. I have nice data on showing major differences by using different growth conditions. Genome sequences has its problems too, as you partly mentioned the discrepancy of your flagellation/motility. So including the sequence is great if you can but not everyone can do it.
Keep thinking what is important to characterize a novel taxa. Sure the more different data one can produce and include is desirable, but some of the minimal descriptions published are a little too ambitious.
What is more important, to show that there is a novel taxa with a minimal description or to have all the different data described the different peoples/groups are interested in and if you can’t do it, not have the new taxa described. Description which are not validated by IJSEM are a waste and cluttering the literature
Juegen,
Thanks for the comment. I should point out that this piece was written by myself, as a scientist in Jonathan Eisen’s lab, and may not reflect his opinions. I agree with most of your comment, but am curious about the following statement “Description which are not validated by IJSEM are a waste and cluttering the literature”. Is it really a waste of time? Those descriptions still provide value to people interested in the taxa in question. Also, if there is ever to be a set of alternative standards would that not have to take place outside IJSEM?