For many years I have been worried about how space travel will affect microbiomes – of the space vehicles and of the residents (people, other animals, plants, etc). This is one of the reasons we started Project MERCCURI and get involved in looking at the microbes on the International Space Station. It is also why we have been collecting papers about microbes on space vehicles (or stations) for a few years (e.g., see the papers tagged space in our Mendeley collection). And we have certainly been writing more and more about space and microbes here. For example see:
- Posts tagged “space”
- Nice series of papers on microbial ecology and space travel
- Antimicrobial coating on a commercial polymer…IN SPACE
- NASA report of interest (from 2012): Genetic inventory (of spacecraft) task final report
- Don’t forget – keep your space robots clean, very clean
- Update on Project MERCCURI: 44/48 samples confirmed for space!
And many many more.
Thus I am both a bit ashamed (because I had not thought deeply about it) and excited to see this news story in GIGAOM: Heading to Mars? Think about packing microbes before food and building supplies bu Signe Brewster and the open access paper behind it: Towards synthetic biological approaches to resource utilization on space missions by Amor A. Menezes, John Cumbers, John A. Hogan and Adam P. Arkin. This paper is a long long read but so far it is fascinating.
Some of their key points are summarized in the following tables and figures from their paper:
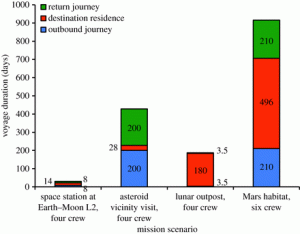
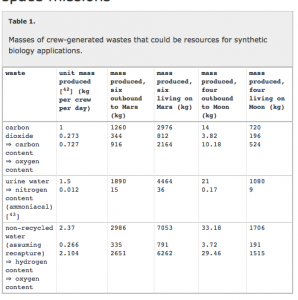
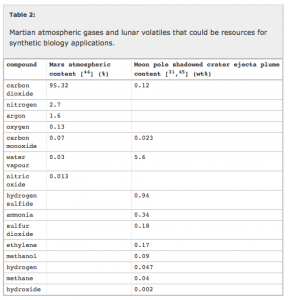
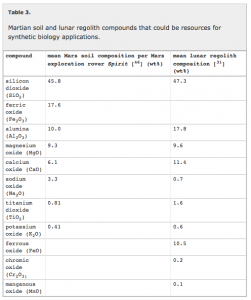
They go through in lots of detail some of the various issues with solving some of the challenges outlined in these tables and the figure. And they conclude that synthetic approaches, such as those involving microorganisms (and also 3D printing) can possible help solve many of these issues.
They even list candidate organisms that have functions that could be of great value:
- Acetobacterium woodii
- strain KN-15 of Methanobacterium
- M. thermoautotrophicum
- deep marine subsurface archaea and bacteria that produce ethane and propane
- GAO
- methanogenic bacteria and cyanobacteria that produce ethane
- Synechocystis sp. PCC 6803 with the efe gene from Pseudomonas syringae pv. Phaseolicola
- A. platensis
- A. maxima
- Fusarium venenatum A3/5
- Synechococcus sp. PCC7002 using the complementation of a cyanobacterial recA null mutation with the E. coli recA gene on a plasmid expression system.
- C. necator H16, also known as R. eutropha H16
- Synechocystis sp. PCC 6803 with the PHA biosynthetic genes of C. necator or with overexpressed sigE
- Synechocystis sp. PCC 6803 with the 4ABH and nhoA genes.
It will take me a while to go through all of this in detail but this is a fascinating article that changes my perspective on “space” and “microbes”. Personally, I do not know much about this area of using microbes as a tool in space travel but at least from my point of view, this paper appears to be pretty groundbreaking and visionary. And it is worth a look.
I am including some responses from Twitter here:
Another response from Twitter
That’s just awesome. We’ve been focusing all this time on keeping microbes OUT of space when we should have been finding a way to use them IN space. It’s amazing how much cargo they’d be saving! Go microbes!